Preface for Supervised Learning#
We already learned how to do linear and logistic regression for descriptive analytics, where the goal is to describe the relationship between some quantity of interest (numerical value like sales, or probability of some class like buy/not buy) and different predictor variables. This is a very important skill!
Now we will instead focus on predictive analytics, where the goal is to make accurate predictions on as-yet unobserved individuals. For all models that we discuss (in this unit and beyond), you are expected to know:
how each model “works” at a fundamental level (and how to describe this to a layperson)
what assumptions (if any) each model makes (i.e., under what conditions should the model be “good”)
what types of relationships are well-suited for a model and which types are not
what parameters we can “tweak” in a model to make it potentially perform better
how the choice of parameters affects the bias-variance tradeoff of the model
Key terms in Predictive Analytics#
The four most important key terms in predictive analytics are:
Generalization error: the error (in the long run) made by a model on individuals it has not yet seen. This quantity cannot be directly observed because it requires predicting on an infinite number of new individuals, but it can be calculated on a specific set of new data. A formula exists that gives the expected generalization error that we will spend a great deal of time discussing (related to the bias-variance tradeoff) in a later unit.
Training data: a set of individuals that are used to “build” the model, i.e., determine its form, estimate coefficients, etc.
Holdout sample: a set of individuals that are used to “test” the model. The model does not look at these individuals when its form is determined, so they are “new” for all intents are purposes.
Overfitting: An overfit model fits the collected data well, but poorly predicts the characteristics of new individuals (biggest fear of a predictive model!) A model is overfit when it becomes “too complex” and includes features that are unique to the particular set of individuals that happen to be in the training data. Since these features are not present in the population at large, we say model is “fitting noise”. An overfit model’s generalization error will be larger than a simpler model that just captured “the gist” of the relationships.
Terminology: response vs. predictors#
When talking about classification and regression models, we use letters to represent what quantity is being predicted and what quantities are used to make the predictions. Sometimes people refer to these quantities as the “response” and “features”.
\(y\) is generally used to represent the quantity that we want to predict (“Junk mail” vs. “Safe mail”, lifetime value, etc.).
\(x\) is generally used to represent the set of quantities that we use to make those predictions (e.g., letter/word frequencies, graduation year, major, etc.)
Usually multiple predictors are used, and \(x_1\), \(x_2\), etc., are used to refer to them.
Response: this is our y variable and the quantity we wish to predict. With classification, \(y\) is a factor with two or more levels. With regression, \(y\) is a number.
Feature: this is a characteristic of an individual, i.e., an x variable, that we will use to make predictions (aka predictor variable).
Regression overview#
Note
Given a set of training data and the responses (\(y\) values) of those individuals, what features and characteristics of these individuals provide information on that individual’s value of \(y\)? How should this information be synthesized and combined into a set of rules that assigns a class to a new individual?
Regression involves predicting some numerical value for an individual, like lifetime, donation amount, purchase quantity) is another.
What is the expected lifetime donation amount for a UT alumnus who graduated in 1964 with an Economics major?
What is the expected monetary value of all claims submitted to an insurance company for a student at UT who has had two prior speeding tickets and drives a Camry?
What is the expected selling price of a house in the 37919 zip code with 4000 square feet, 2.5 baths, and a pool?
Note
In data mining, we use the word regression to describe any technique that predicts a numerical value, regardless of whether “linear regression” is used to do so!
In fact, the vanilla partition model (decision tree model) was first called CART for Classification And Regression Trees, even though numerical predictions are made by a completely different process than a linear regression.
There are many ways to approach modeling a numerical value. Take the UT alumni dataset again where the following characteristics are known about graduates:
Academic information (date of graduation, college, major, degree, number of degrees)
Student life (going Greek, student government, etc.)
Familial relationships (did spouse or children attend UT)
Donation history (total so far, total number of gifts, actual amount the last few years)
How can we convert this information to make predictions about an alumnus’ lifetime donation amount?
If \(GradYear < 1960\) and \(College=Business\) and \(Greek=TRUE\), then predict $12,000.
\(Amount = 82 - 0.002(GradYear) + 3005(NumDegrees) + 1.5(SizeFirstGift)\)
etc
Classification overview#
The task of classification is perhaps the most popular pillar of data mining.
Note
Given a set of training data and the class labels of those individuals, what features and characteristics of these individuals provide information on that individual’s class? How should this information be synthesized and combined into a set of rules that assigns a class to a new individual?
Emails have two class labels: junk (spam) or legitimate. What are the characteristics of junk vs. legitimate email? How can an algorithm determine whether an email is junk or not?
Alumni have a few class labels: never donors (those who will never donate to UT), major donors (those who donate more than $10,000 over their life), and casual donors (the rest). What characterizes each class? How can an algorithm decide who is who?
Imagine that the development office at UT has provided you with a large set of data on previous graduates:
Academic information (date of graduation, college, major, degree, number of degrees)
Student life (going Greek, student government, etc.)
Familial relationships (did spouse or children attend UT)
Donation history (total so far, total number of gifts, actual amount the last few years)
Can we leverage this information in such a way that it accurately classifies alumni as never donors, casual donors, and major donors?
if \(GradYear < 1960\) and \(College=Business\) and \(Greek=TRUE\), then major donor?
\(P(never) = e^{-5 -0.02GradYear + 3.1NumDegrees + 0.00043(SizeFirstGift)}\). If \(P(never) > 0.5\), then never donate?
Many business questions are classification problems.
Classify customers as a future buyer or non-buyer.
Classify a web surfer as one who will click on an ad or not.
Classify political leaning: left, middle, right.
Classify outcome of a new restaurant: succeed, fail.
Classify customer’s future loan payment status: on time, late, default.
Classify loyalty of customers: churn vs. stay.
What other problems might businesses face that can be treated as a classification problem?
Algorithms for predictive analytics#
There are many data mining algorithms (sets of rules that convert the information available about each individual into a predicted class) for classification:
Linear (vanilla and regularized) regression: (predicts numerical response)
Logistic (vanilla and regularized) regression: two class problem (predicts probabilities)
Multinomial logistic regression: multi-class problem (predicts probabilities)
Naive Bayes Classifiers: multi-class problem (predicts probabilities/likelihood scores)
Decision trees (vanilla partition model, random forests, boosted trees, etc.): numerical responses, multi-class classification problems
Nearest Neighbors: numerical responses, multi-class classification problems
Neural Networks: numerical responses, multi-class classification problems
Support Vector Machines: numerical responses, two-class classification problems
Numerical response: 4.51, 871.33, etc. Two-class problem: “yes” vs. “no”. Multi-class problem: “red”, “yellow”, “green”, “blue”.
Evaluation Metrics#
The terms and quantities used to evaluate classification models vs. regression models are different.
Note
Regression
RMSE: the “root mean squared error” (typical size of error made by model)
\(\mathbf{R^2}\): the “R-squared”
MAE: mean absolute error, etc.
Classification
Misclassification Rate
Accuracy
AUC (area under the ROC curve), false positive rate, false negative rate, Kappa, etc.
Note
Metrics for descriptive vs. predictive analytics
When doing descriptive analytics, we were interested in finding a model that provided a reasonable reflection of reality since we wanted to talk about what relationships “looked like”. In that context, we assessed models by discussing their RMSE, \(R^2\), or misclassification rate and we compared models using the AIC (“distance” from “the truth” to the model) on the training data.
Note
We don’t really care how the models we use in predictive analytics fit the training data! To evaluate the predictive performance of models, we’ll judge models based on how well they “generalize” to new data. A “good” model will have a “small” error when making predictions about individuals that were not used to determine the model’s form.
Generalization Error (GE)#
When doing predictive analytics, special treatment of the collected data is necessary:
Training Data - used for model-fitting, exploration, and comparison
Holdout Sample - used only once to confirm predicted accuracy of final selected model.
When doing predictive analytics, the collected data is usually randomly split into the training data and the holdout sample. A final model is selected using the training data, and a “sanity check” is performed by making predictions on the holdout sample to make sure the model is making errors with roughly the same size or frequency as expected.
When evaluating a predictive model and choosing between them, how well the model fits the training data (the set of individuals used to build the model) is more or less irrelevant. We already know the answers to these individuals anyway!
Instead, we want to choose the model that has the lowest generalization error (accuracy, AUC, RMSE, etc.), i.e., in the long run, makes the most accurate predictions on new (current or future) individuals the model has not seen.
The multiple linear regression model (predicting response rate for a census block group from all predictors and all two-way interactions) fits the training data well (typical error 3.5) but doesn’t generalize well (typical error 5.9 on holdout). This model is “overfit”.
library(tidymodels)
library(regclass)
data(CENSUS)
Show code cell output
── Attaching packages ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────── tidymodels 1.3.0 ──
✔ broom 1.0.7 ✔ recipes 1.1.1
✔ dials 1.4.0 ✔ rsample 1.2.1
✔ dplyr 1.1.4 ✔ tibble 3.2.1
✔ ggplot2 3.5.1 ✔ tidyr 1.3.1
✔ infer 1.0.7 ✔ tune 1.3.0
✔ modeldata 1.4.0 ✔ workflows 1.2.0
✔ parsnip 1.3.0 ✔ workflowsets 1.1.0
✔ purrr 1.0.4 ✔ yardstick 1.3.2
── Conflicts ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────── tidymodels_conflicts() ──
✖ purrr::%||%() masks base::%||%()
✖ purrr::discard() masks scales::discard()
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
✖ recipes::step() masks stats::step()
Loading required package: bestglm
Loading required package: leaps
Loading required package: VGAM
Loading required package: stats4
Loading required package: splines
Attaching package: ‘VGAM’
The following object is masked from ‘package:workflows’:
update_formula
Loading required package: rpart
Attaching package: ‘rpart’
The following object is masked from ‘package:dials’:
prune
Loading required package: randomForest
randomForest 4.7-1.2
Type rfNews() to see new features/changes/bug fixes.
Attaching package: ‘randomForest’
The following object is masked from ‘package:ggplot2’:
margin
The following object is masked from ‘package:dplyr’:
combine
Important regclass change from 1.3:
All functions that had a . in the name now have an _
all.correlations -> all_correlations, cor.demo -> cor_demo, etc.
#Split data into a training set and holdout sample
set.seed(474)
tt_split <- initial_split(CENSUS, prop = 0.7)
TRAIN <- training(tt_split)
HOLDOUT <- testing(tt_split)
#Fit model on TRAINING data (not immediately interesting)
REC <- recipe(ResponseRate ~ ., TRAIN) %>%
step_nzv(all_predictors()) %>%
step_corr(all_predictors()) %>%
step_lincomb(all_predictors()) %>%
step_interact(terms = ~ all_predictors()^2) %>%
step_pca(all_predictors(), num_comp = 50)
MODEL <- workflow() %>%
add_recipe(REC) %>%
add_model(linear_reg()) %>%
fit(data = TRAIN)
#typical error on train
TRAIN %>%
select(ResponseRate) %>%
bind_cols(predict(MODEL, new_data = TRAIN)) %>%
metrics(truth = 'ResponseRate', estimate = '.pred')
#typical error on holdout
HOLDOUT %>%
select(ResponseRate) %>%
bind_cols(predict(MODEL, new_data = HOLDOUT)) %>%
metrics(truth = 'ResponseRate', estimate = '.pred')
| .metric | .estimator | .estimate |
|---|---|---|
| <chr> | <chr> | <dbl> |
| rmse | standard | 4.8815313 |
| rsq | standard | 0.5723798 |
| mae | standard | 3.7331225 |
| .metric | .estimator | .estimate |
|---|---|---|
| <chr> | <chr> | <dbl> |
| rmse | standard | 5.4099633 |
| rsq | standard | 0.4943867 |
| mae | standard | 3.9593914 |
Note
The abstract nature of the generalization error
The generalization error of a model is the average error made by the model on “new” individuals in the long run (i.e., after thousands, if not millions, of predictions are made on as-yet-unseen individuals).
We can never directly observe this value; we can compute this error on a particular holdout sample (or sample of new individuals).
The error on a particular set of new individuals in a holdout sample is a random quantity (dependent on what new individuals happen to be in it). It is possible that the “right” model could predict poorly on an odd collection of individuals!
Note
The best we can do is to somehow estimate a model’s generalization error, keeping in mind that our estimate is imperfect, and that we are rarely compelled to choose the model with the lowest estimated generalization error.
There are many ways to estimate the generalization error of a model, but we’ll stick with \(K\)-fold crossvalidation.
\(K\)-fold crossvalidation and One Standard Deviation Rule#
\(K\)-fold cross-validation#
We will estimate the GE of a model with the \(K\)-fold cross-validation procedure:
Randomly split up the training data into \(K\) equal sized chunks.
In round one of the procedure, the last chunk of the training data is used as a validation (“pseudo-holdout”) sample. The model is fit using the remaining \(K-1\) chunks of the training data, and the typical error on the pseudo-holdout set is recorded.
In round two, the second to last chunk of the training data is used as the pseudo-holdout. The model is fit using the remaining \(K-1\) chunks of the training data, and the typical error on the pseudo-holdout set is recorded.
After all \(K\) chunks have been used as the pseudo-holdout set once, the estimated generalization error is the average of these \(K\) errors.
This estimated generalization error is typically an underestimate of what the actual error on brand-new individuals will be.
The value of \(K\) is a choice (usually 5 or 10). When \(K\) is small, the algorithm goes faster but the generalization error is typically underestimated by more. When \(K\) is large, the algorithm goes slower but the GE is underestimated by less.
set.seed(474)
tt_split <- initial_split(CENSUS, prop = 0.7)
TRAIN <- training(tt_split)
HOLDOUT <- testing(tt_split)
REC <- recipe(ResponseRate ~ ., TRAIN) %>%
step_nzv(all_predictors()) %>%
step_corr(all_predictors()) %>%
step_lincomb(all_predictors()) %>%
step_interact(terms = ~ all_predictors()^2) %>%
step_pca(all_predictors(), num_comp=50)
WF <- workflow() %>%
add_recipe(REC) %>%
add_model(linear_reg())
RES <- WF %>%
fit_resamples(vfold_cv(TRAIN, v = 5), metrics = metric_set(rmse, rsq, mae))
collect_metrics(RES)
| .metric | .estimator | mean | n | std_err | .config |
|---|---|---|---|---|---|
| <chr> | <chr> | <dbl> | <int> | <dbl> | <chr> |
| mae | standard | 3.820156 | 5 | 0.06253162 | Preprocessor1_Model1 |
| rmse | standard | 5.016267 | 5 | 0.09437582 | Preprocessor1_Model1 |
| rsq | standard | 0.548714 | 5 | 0.01076380 | Preprocessor1_Model1 |
Important
Compare the cross validation results with the previous TRAIN/HOLDOUT results.
Illustration if \(K=4\) (holdout sample not displayed).
par(mfrow=c(2,2))
par(mar=c(1,1,1,1))
K <- 4
for (k in 1:K) {
plot(0,0,type="n",axes=FALSE,xlim=c(0,1),ylim=c(0,1),xlab= "",ylab= "",main=paste("Round",k))
rect(0,0,1,1)
rect( 1-k/K,0,1-k/K+1/K,1, col=grey(.8))
#for (i in 1:K) { lines( c(i/K,i/K),c(0,1) ) }
label.loc <- 1- (1:K)/K + 1/(2*K)
text( label.loc[k], 0.5, "Validation")
for (z in setdiff(1:K,k)) { text( label.loc[z], 0.5, "Training") }
}
par(mfrow=c(1,1))
par(mar=c(5, 4, 4, 2) + 0.1)
If time allows, the \(K\)-fold crossvalidation procedure is repeated with a different random assignment of individuals into \(K\) chunks. Ultimately, the final estimate of the generalization error is the average of the estimates found in each repeat. Repeating the procedure helps to guard against chance assignments into chunks that might yield a particularly low or high estimated error.
No guarantee that lower estimate = better model#
The estimated generalization error that emerges from \(K\)-fold cross-validation is truly only an estimate of the error that model will make on new data.
Since the procedure incorporates randomness to estimate the error, the number that emerges
Since a particular set of new individuals invariably will have their own set of quirks, there’s a chance that a great model happens to predict poorly on them, and there’s a chance a poor model happens to predict well.
Even the “correct” model might not predict all that great on a small set of new individuals. For example, a correct model for a fair coin would predict half of new flips to be heads and half to be tails, but the next 8 flips could all be heads!
Note
After trying out many models, the one with the lowest estimated generalization error will rarely be the one that actually performs the best on a holdout sample or on a particular set of as-yet unseen individuals. There is no compelling reason to always choose this model if other comparable models have more appealing properties (e.g., simpler, easier to explain).
A few hundred models to predict the body fat percentage of individuals were built on a training sample. The estimated generalization errors were found with \(5\)-fold cross validation. The plot shows this estimate (horizontal axis) versus the actual generalization error on a particular holdout sample (vertical axis). A diagonal line is drawn where the two errors are equal. Each dot represents a model.
The actual generalization errors of models is typically above the estimated generalization errors. This is typical. Though we do our best, we typically underestimate the amount of error a model will make on new data.
The model with the lowest estimated generalization error (dot on far left) is nowhere close to the model that ended up predicting the best on the holdout sample.
One standard deviation rule#
If we have fit many different kinds of models, the one standard deviation rule provides some guidance over which models are acceptable.
Note
If two models have estimated generalization errors within one standard deviation of each other, there is no compelling statistical reason to prefer one vs. the other.
What does this “one standard deviation” refer to? During repeated \(K\)-fold cross-validation (say 10 repeats were performed), we have 10 estimates of the generalization error. The standard deviation of these 10 estimates provides a reasonable “buffer of equivalence”, meaning that two models with estimated errors within one standard deviation of each other are more or less equivalent. If no repeats were done, it is the standard deviation of the \(K\) estimates instead.
The final estimate of the (RMSE) generalization error was found to be 0.8792 (average of the 10 estimates from repeated \(K\)-fold crossvalidation). The standard deviation of these estimates is 0.024. Thus, any model whose estimated generalization error is 0.903 or less, for all intents are purposes, generalizes “just as well” as this one.
#estimated generalization errors from 10 repeats of K-fold crossvalidation
estimates <- c(.871, .889, .834, .911, .891, .865, .899, .861, .867, .904)
#average of estimates
mean(estimates)
#standard deviation of estimates
sd(estimates)
#one standard deviation rule
mean(estimates) + sd(estimates)
Two different models yielded the following estimated generalization errors during 10 repeats of 5-fold crossvalidation. Is one preferred vs. the other?
set.seed(474); model1 <- round( runif(10,.01,.05), digits = 3)
set.seed(475); model2 <- round( runif(10,.02,.08), digits = 3)
model1 #estimated errors of K-fold cross validation over 10 repeats
## [1] 0.023 0.037 0.030 0.022 0.025 0.047 0.027 0.031 0.012 0.038
model2 #estimated errors of K-fold cross validation over 10 repeats
## [1] 0.070 0.052 0.035 0.049 0.042 0.045 0.064 0.022 0.060 0.059
mean(model1); sd(model1) #final estimate and its standard deviation
## [1] 0.0292
## [1] 0.00981835
mean(model2); sd(model2) #final estimate and its standard deviation
## [1] 0.0498
## [1] 0.01445145
#divide by larger of 2 SDs
(mean(model1) - mean(model2) ) / max(c(sd(model1),sd(model2)))
## [1] -1.425462
The estimated generalization error of the model 1 is over 2 standard deviations BELOW the estimated generalization error of model 2. There IS compelling evidence to believe that model 1 is better (though surely there are some sets of a small number of new individuals that model 2 will outperform model 1; model 1 is likely the better choice in the long run).
Important
Now compare the following two models.
Is one model better than the other?
If yes, which one?
If no, why?
set.seed(474)
tt_split <- initial_split(CENSUS, prop = 0.7)
TRAIN <- training(tt_split)
HOLDOUT <- testing(tt_split)
REC <- recipe(ResponseRate ~ ., TRAIN) %>%
step_nzv(all_predictors()) %>%
step_corr(all_predictors()) %>%
step_lincomb(all_predictors()) %>%
step_interact(terms = ~ all_predictors()^2) %>%
step_pca(all_predictors(), num_comp=tune())
WF <- workflow() %>%
add_recipe(REC) %>%
add_model(linear_reg())
# model with 50 principal components
RES <- WF %>%
finalize_workflow(list(num_comp=50)) %>%
fit_resamples(vfold_cv(TRAIN, v = 5), metrics = metric_set(rmse, rsq, mae))
collect_metrics(RES) %>%
select(.metric, mean, std_err)
| .metric | mean | std_err |
|---|---|---|
| <chr> | <dbl> | <dbl> |
| mae | 3.820156 | 0.06253162 |
| rmse | 5.016267 | 0.09437582 |
| rsq | 0.548714 | 0.01076380 |
# model with 25 principal components
RES <- WF %>%
finalize_workflow(list(num_comp=25)) %>%
fit_resamples(vfold_cv(TRAIN, v = 5), metrics = metric_set(rmse, rsq, mae))
collect_metrics(RES) %>%
select(.metric, mean, std_err)
| .metric | mean | std_err |
|---|---|---|
| <chr> | <dbl> | <dbl> |
| mae | 3.8543169 | 0.09056184 |
| rmse | 5.0424506 | 0.13432533 |
| rsq | 0.5435684 | 0.01718454 |
Occam’s Razor#
The heuristic of “Occam’s razor” is quite popular.
``Given two explanations for the same thing, the simpler one is usually the correct one”.
Now since “all models are wrong; some are useful”, the notion of being “correct” gets thrown out the window, so Occam’s razor simply suggests choosing the simplest model whose estimated generalization error is no more than 1 standard deviation above the lowest error.
Generally, it’s better to go with a noticeably simpler model only if it’s performance just a little worse (much less than 1 SD) than a more complex model.
Strategy for Model Selection#
In the example above, we compared two parameter values for num_comp. How to automate such comparison for models with more parameter valuess? We can use the tune_grid function:
set.seed(474)
tt_split <- initial_split(CENSUS, prop = 0.7)
TRAIN <- training(tt_split)
HOLDOUT <- testing(tt_split)
REC <- recipe(ResponseRate ~ ., TRAIN) %>%
step_nzv(all_predictors()) %>%
step_corr(all_predictors()) %>%
step_lincomb(all_predictors()) %>%
step_interact(terms = ~ all_predictors()^2) %>%
step_pca(all_predictors(), num_comp=tune())
WF <- workflow() %>%
add_recipe(REC) %>%
add_model(linear_reg())
RES <- WF %>%
tune_grid(
grid = expand.grid(
num_comp = c(25, 30, 35, 40, 45, 50)
),
resamples = vfold_cv(TRAIN, v = 5)
)
collect_metrics(RES)
| num_comp | .metric | .estimator | mean | n | std_err | .config |
|---|---|---|---|---|---|---|
| <dbl> | <chr> | <chr> | <dbl> | <int> | <dbl> | <chr> |
| 25 | rmse | standard | 5.0412541 | 5 | 0.09221150 | Preprocessor1_Model1 |
| 25 | rsq | standard | 0.5440121 | 5 | 0.01418786 | Preprocessor1_Model1 |
| 30 | rmse | standard | 5.0340176 | 5 | 0.08750424 | Preprocessor2_Model1 |
| 30 | rsq | standard | 0.5450178 | 5 | 0.01326056 | Preprocessor2_Model1 |
| 35 | rmse | standard | 5.0362908 | 5 | 0.09536618 | Preprocessor3_Model1 |
| 35 | rsq | standard | 0.5445897 | 5 | 0.01175674 | Preprocessor3_Model1 |
| 40 | rmse | standard | 5.0265969 | 5 | 0.09507688 | Preprocessor4_Model1 |
| 40 | rsq | standard | 0.5465194 | 5 | 0.01232073 | Preprocessor4_Model1 |
| 45 | rmse | standard | 5.0321313 | 5 | 0.09448088 | Preprocessor5_Model1 |
| 45 | rsq | standard | 0.5455938 | 5 | 0.01145398 | Preprocessor5_Model1 |
| 50 | rmse | standard | 5.0162670 | 5 | 0.09437582 | Preprocessor6_Model1 |
| 50 | rsq | standard | 0.5487140 | 5 | 0.01076380 | Preprocessor6_Model1 |
Important
Does the choice of parameter values improve the modeling performance? Use the one-standard-deviation rule!
The no free lunch theorem states that there is no one type of model that works best for every problem.
All models are (over) simplified representations of reality (simplifications are almost always necessary to make the problem under study tractable).
For a simplified representation to be “good”, certain assumptions must be true.
When assumptions hold true, the model is “good” and provides a reasonable reflection of reality, so it should work “well” for the task at hand.
Issue: assumptions are of course not always true! A model may do quite well in some situations, but it can do terribly in others (e.g., linear regression failing to capture a nonlinear relationship).
Example: How do you calculate the time it takes a chicken to fall to the ground. Physicist will “assume a spherical chicken in a vacuum”.
Note
In practice, when tasked with predictive analytics problem, you must try out many algorithms and dozens if not hundreds of particular models. The performance of each must be evaluated, and the most suitable algorithm and model for that specific problem needs to be identified.
While it’s ok to have a “go-to” model (mine is the random forest), the no free lunch theorem tells us that we must explore a variety a models to find the “best” one. There’s a lot of trial and error in predictive analytics.
To estimate generalization error of a model, we need to check to see how well it predicts on new data. But, unless new data is expected to come in very soon, we are stuck with the data currently on hand! What do we do?
Split the original data into a training sample and a holdout sample (also called the test set).
Estimate the generalization errors of many models using the training data via repeated \(K\)-fold crossvalidation.
Identify the subset of models that have estimated generalization errors within one standard deviation of the lowest.
Of those, choose your favorite: the overall lowest, the easiest to interpret to explain to your boss, the simplest model, etc.
Using the selected model, find the actual generalization error for your holdout sample. Compare this to the estimated generalization error.
A “good” model (not overfit) should not have the error on the holdout sample be much larger than the estimated error during \(K\)-fold crossvalidation (“much larger” is in the eye of the beholder, maybe 10% or so, though I’ve said up to 25% in the past).
Note
Don’t select a model using the holdout sample
The holdout sample is used exactly once, and that is when the final selected model undergoes evaluation. Its purpose is solely to verify that the model’s error on a set of new data is about what the estimated generalization error suggested (it’s ok to be maybe 10% or so higher).
If, out of curiosity, you make predictions on the holdout sample using a different model, and find its error is lower than the selected model, you don’t switch models.
The holdout sample is never used to choose between different models.
One model having a lower error on a particular holdout sample could very well just be “good luck”.
Overfitting#
A model is overfit when it predicts well on the individuals in the training data but poorly on new individuals it has not seen.
Overfitting occurs when a model is “too complex”. Adding additional complexities “beyond what is needed” results in the model incorporating quirks that are unique to the individuals in that set of training data. When an overfit model makes predictions on new individuals, they will be worse than had the model captured just the essence of the relationships and remained simpler.
The \(overfit\_demo\) in \(regclass\) has some interesting examples.
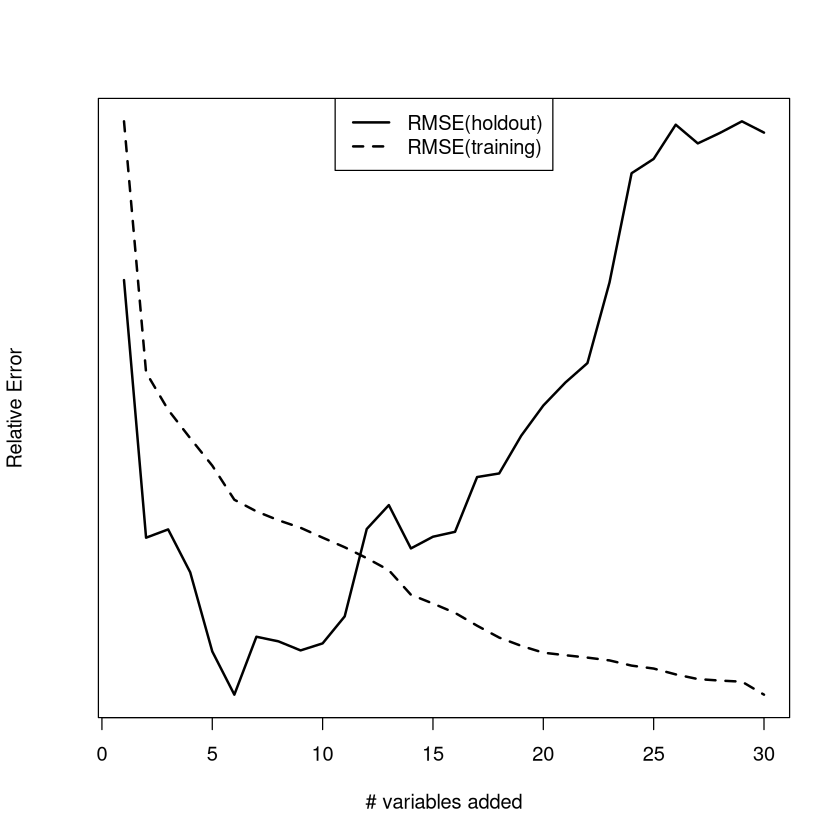
Overfitting demo (one of top 5 conceptual examples in the course)#
The typical errors of a linear regression model on the training data and a holdout sample (of new individuals) is considered. As the number of variables increases, the performance on the training data steadily improves (values of the RMSE get smaller) while the typical size of the error on the holdout sample (RMSE) gets larger.
Avoiding overfitting#
Obviously, we want to avoid ending up with a model that is overfit. How do we avoid doing so?
Note
It is impossible to determine whether or not a model is overfit by looking at its performance on the training data. By looking at the estimated generalization error with \(K\)-fold crossvalidation, and by selecting a model with an appropriate low expected generalization error (within the buffer given by the one standard deviation rule), we guard against overfitting.
Note: more complex models have a better chance of being overfit than simpler models. If two models are really close to each other in terms of performance, go with the simpler one!
Closing remarks#
Choosing the right modeling approach is important. Choosing the right complexity of the right modeling approach is just as important.
A model that is too “simple” may be relatively insensitive to the particular set of values recorded in the training sample (low variance), but it may miss out on key features of relationships that make its performance suffer (high bias). In other words, the model is underfit.
A model that is too “complex” may fit the training data fantastically (low bias). However, because the form of the model is quite sensitive to the particular values in the training data, it suffers from high variance and may not generalize well to new data. In other words, the model is overfit.
Because we can’t just plug in values into the equation that gives us the expected generalization error, we have to find the optimal complexity of a model (best balance between bias and variance) with crossvalidation techniques. The mechanics of searching for a “good” model, tuning parameters, and estimating error can be quite computationally intensive, but it pays off in the end!
We have learned and will continue to learn about phenomenally sophisticated, cutting edge models. The no free lunch theorem guarantees that these are not always necessarily the better choice (though they often are). Do not take this for granted!
Note
In business analytics, marketing, and finance, surprisingly simple models sometimes predict more accurately than more complex, sophisticated models. Don’t use a bulldozer when a shovel will do.
Upshot: although it “feels weird” to select as your final model one that doesn’t fit the training data as well as other models you have considered (i.e., it has a higher bias), often that is the right thing to do. If the model-building procedure has a lower variance than other procedures, it might just have a lower generalization error.