library(tidymodels)
library(neuralnet)
Show code cell output
── Attaching packages ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────── tidymodels 1.3.0 ──
✔ broom 1.0.7 ✔ recipes 1.1.1
✔ dials 1.4.0 ✔ rsample 1.2.1
✔ dplyr 1.1.4 ✔ tibble 3.2.1
✔ ggplot2 3.5.1 ✔ tidyr 1.3.1
✔ infer 1.0.7 ✔ tune 1.3.0
✔ modeldata 1.4.0 ✔ workflows 1.2.0
✔ parsnip 1.3.0 ✔ workflowsets 1.1.0
✔ purrr 1.0.4 ✔ yardstick 1.3.2
── Conflicts ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────── tidymodels_conflicts() ──
✖ purrr::%||%() masks base::%||%()
✖ purrr::discard() masks scales::discard()
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
✖ recipes::step() masks stats::step()
Attaching package: ‘neuralnet’
The following object is masked from ‘package:dplyr’:
compute
Neural Networks and Deep Learning#
Neural network models emerged from early attempts to model how neurons in the brain might work. While they have had only limited success in actually modeling anything biological and their popularity has ebbed and flowed (mostly due to computational limitations over time), they have become quite useful in machine learning.
In fact, you may have heard of the phrase ``deep learning”, which is used to solve problems in speed recognition, imagine recognition, 3-D object recognition. Deep learning is just a very large, very complex neural network.
So how does it work?
In 1958, Frank Rosenblatt developed the perceptron (in a sense is the most primitive neural network) to make classifications.
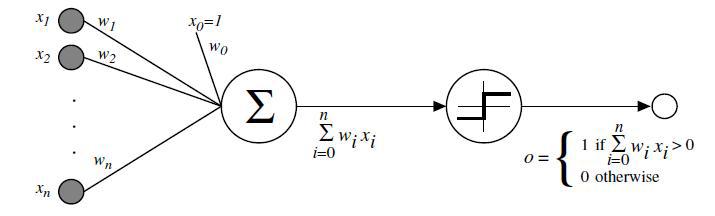
The perceptron takes input \(x_1\), \(x_2\), etc. (the values of predictor variables) and calculates the weighted sum \(w_0 + w_1 x_1 + w_2 x_2 + \ldots\), where weights can be positive or negative. However, it outputs either a 0 or a 1, depending on whether the value of the weighted sum is positive or negative.
If you think the process of making a prediction with the perceptron sounds like linear regression, you’re right.
In fact, the perceptron looks a lot like a linear regression model trying to predict the probability of a level and then classifying accordingly. Linear regression is usually a pretty bad model for this application (which is why we use logistic regression instead).
Activation Function#
People realized that instead of transforming the weighted sum of predictors to an output using a ``step function” (0 if less than some threshold, 1 if greater), a better idea would be to feed the weighted sum into a more general activation function. One popular choice is the logistic function.
When the perceptron uses the logistic activation function, the end result is logistic regression, which we can use the model the probability an individual possesses one of two classes.
By taking the weighted sum of predictor variables and passing it through the logistic activation function, the output becomes a number between 0 and 1 and could be interpreted as a probability. The upshot has been the logistic regression has been re-invented.

There are many choices of activation functions, though by far the most common is the logistic (aka sigmoid) due to some ``nice” theoretical properties.
Show code cell source
par(mfrow=c(2,2))
plot( seq(-4,4,by=0.01), c( rep(0,401),rep(1,400)),type="l",xlab="Weighted Sum",ylab="Output")
legend("topleft","Step")
curve( 1/(1+exp(-x)), from=-4,to=4,xlab="Weighted Sum",ylab="Output")
legend("topleft","Logistic/Sigmoid")
curve( atan(x), from=-4,to=4,xlab="Weighted Sum",ylab="Output")
legend("topleft","ArcTan")
curve( x/(1+abs(x)), from=-4,to=4,xlab="Weighted Sum",ylab="Output")
legend("topleft","Softsign")
par(mfrow=c(1,1))
There’s not a huge variety in what activation functions look like.

Example: MNIST Digits#
How do you personally figure out what digit is what? What features do you look for?

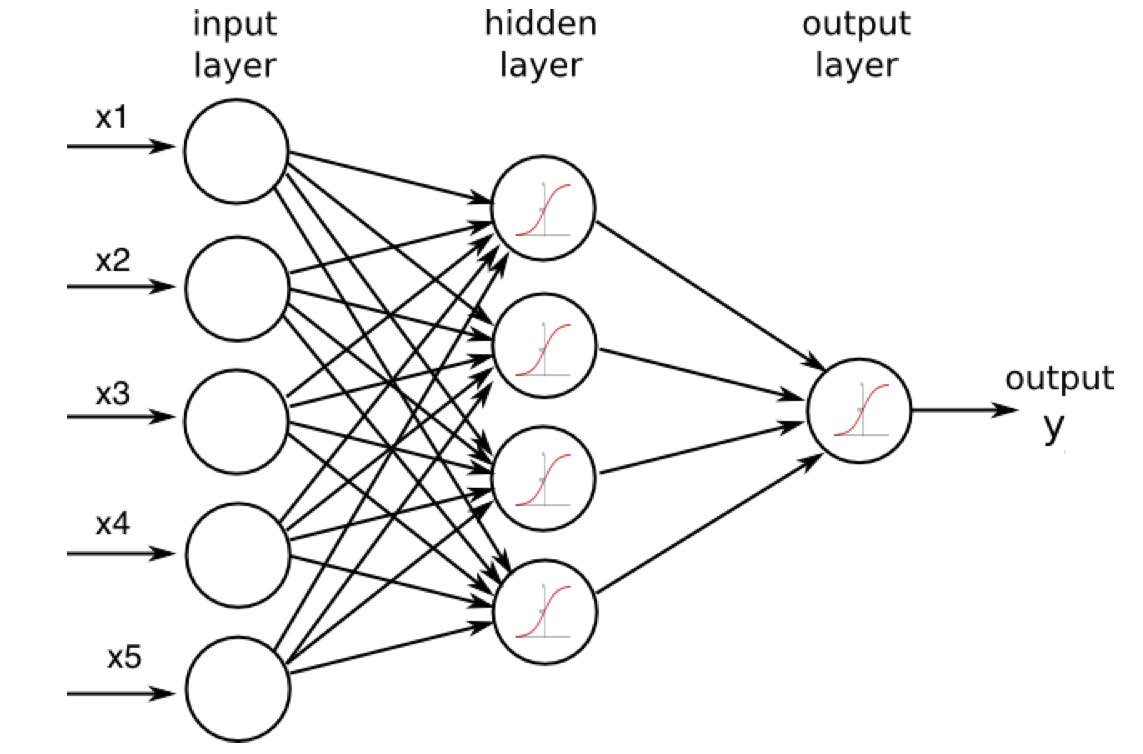
The hidden layer in the neural network might ``construct” features like:
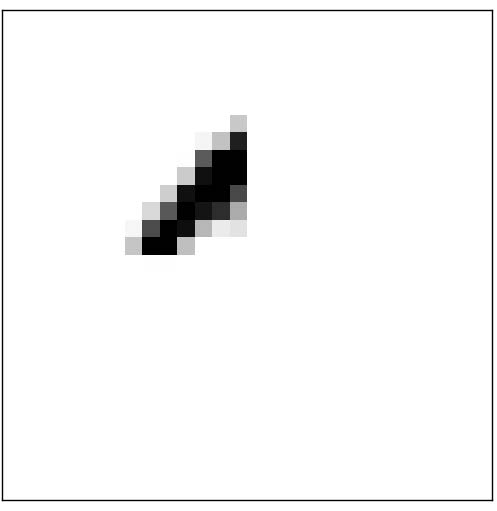
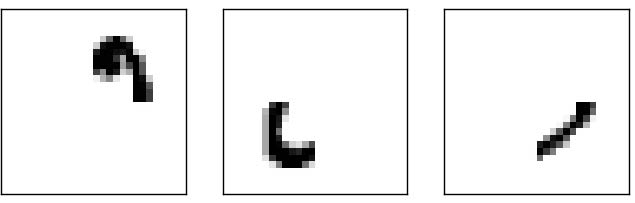


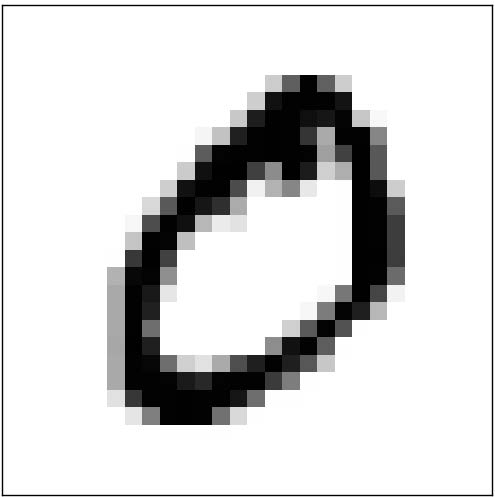
A ``weighted sum” of those four features could easily produce a 0! Other sets of features would be able to produce other digits.

Tuning Parameters#
When building a neural network for predictive modeling, you need to design it:
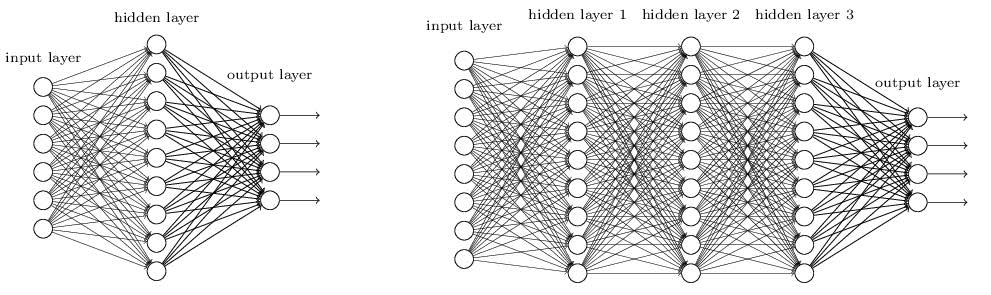
Number of hidden layers: More layers = lower bias (fitting the training data well) but larger variance and larger risk of overfitting unless you have a really big dataset.
Number of neurons in each hidden layer: essentially, the number of new variables to create from the original predictors. More neurons = lower bias but larger variance and larger risk of overfitting.
“Weight decay” (regularization): penalty given to large weights in the weighted sum to prevent overfitting and to improve generalization. This will prevent any particular predictor/feature from contributing “too much” and makes the model somewhat less sensitive to the set of individuals in the training data. Small weights make the modeled relationships “more linear” and prevents crazy curviness that may be unique to the training set. Not all implementations of the algorithm have this parameter.
Example and Pros and Cons#
The \(neuralnet\) lets us explore some aspects of the neural network model (we will not be using it for \(train\)), but the data frame has to have categorical variables converted to indicator variables and each variable needs to be scaled.
data(TIPS, package='regclass')
#Replace categorical variables with indicator variables (using only some of the predictors here)
DATA <- model.matrix(~TipPercentage+Bill+Gender+Smoker+PartySize,data=TIPS)[,-1]
#Create variables to store the mean and standard deviation of the y variable; need those later
mean.y <- mean(TIPS$TipPercentage); sd.y <- sd(TIPS$TipPercentage)
DATA <- scale(DATA) #Scale the data
NNET <- neuralnet(TipPercentage~Bill+GenderMale+SmokerYes+PartySize,data=DATA,
hidden=3,linear.output = TRUE,stepmax=1e6)
The \(hidden\) argument tells it how many neurons in the hidden layer (can make this a vector to add more hidden nodes). \(stepmax\) dictates how many search steps the algorithm is allowed to take to find the optimal set of weights.
Visualizing the neural network with neuralnet#
Warning: you may not be able to knit the plot of a neural network because of some odd choices the authors of the package made with the plot syntax (in fact, there is some chance your computer may not even be able to make the plot at all).

Making predictions with neuralnet#
The writers of the \(neuralnet\) package also decided to make their neural network model incompatible with \(predict\). Instead, you must use \(compute\).
Note: when giving it a dataset on which to make predictions, the columns must match up exactly with the columns used to train the model (unlike every other model we’ve discussed). The example below uses everything but the TipPercentage column (since that is the \(y\) variable and not an \(x\) variable that appears in the predictor matrix).
predictions.nnet <- as.numeric( compute(NNET,DATA[,-1])$net.result ) #ie without column of y
predictions.nnet <- predictions.nnet*sd.y + mean.y #unscale
#RMSE on training data; not the most interesting
sqrt( mean( (TIPS$TipPercentage - predictions.nnet)^2 ) )
Examples with \(tidymodels\)#
Example: neural networks (classification)#
data(EX6.WINE, package='regclass')
# This is not usually right if want to investigate testing performances.
TRAIN <- EX6.WINE
REC <- recipe(Quality ~ ., TRAIN) %>%
step_normalize(all_numeric_predictors()) %>%
step_dummy(all_nominal_predictors()) %>%
step_nzv(all_predictors()) %>%
step_corr(all_predictors()) %>%
step_lincomb(all_predictors())
WF <- workflow() %>%
add_recipe(REC) %>%
add_model(mlp( mode = 'classification', hidden_units = tune(), penalty = tune() ))
GRID <- expand.grid( hidden_units = 1:7, penalty = 10^seq(-2,2,length=20) )
RES <- WF %>%
tune_grid(
resamples = vfold_cv(TRAIN, v = 5),
grid = GRID,
)
METRICS <- collect_metrics(RES)
METRICS
| hidden_units | penalty | .metric | .estimator | mean | n | std_err | .config |
|---|---|---|---|---|---|---|---|
| <int> | <dbl> | <chr> | <chr> | <dbl> | <int> | <dbl> | <chr> |
| 1 | 0.01000000 | accuracy | binary | 0.8188889 | 5 | 0.004771888 | Preprocessor1_Model001 |
| 1 | 0.01000000 | brier_class | binary | 0.1615436 | 5 | 0.002081009 | Preprocessor1_Model001 |
| 1 | 0.01000000 | roc_auc | binary | 0.8960086 | 5 | 0.006585751 | Preprocessor1_Model001 |
| 2 | 0.01000000 | accuracy | binary | 0.8437037 | 5 | 0.007041909 | Preprocessor1_Model002 |
| 2 | 0.01000000 | brier_class | binary | 0.1509120 | 5 | 0.002011134 | Preprocessor1_Model002 |
| 2 | 0.01000000 | roc_auc | binary | 0.9148295 | 5 | 0.005190027 | Preprocessor1_Model002 |
| 3 | 0.01000000 | accuracy | binary | 0.8451852 | 5 | 0.013902463 | Preprocessor1_Model003 |
| 3 | 0.01000000 | brier_class | binary | 0.1482455 | 5 | 0.003201304 | Preprocessor1_Model003 |
| 3 | 0.01000000 | roc_auc | binary | 0.9179203 | 5 | 0.007189008 | Preprocessor1_Model003 |
| 4 | 0.01000000 | accuracy | binary | 0.8388889 | 5 | 0.004721314 | Preprocessor1_Model004 |
| 4 | 0.01000000 | brier_class | binary | 0.1491965 | 5 | 0.001408213 | Preprocessor1_Model004 |
| 4 | 0.01000000 | roc_auc | binary | 0.9150855 | 5 | 0.005121324 | Preprocessor1_Model004 |
| 5 | 0.01000000 | accuracy | binary | 0.8555556 | 5 | 0.007790994 | Preprocessor1_Model005 |
| 5 | 0.01000000 | brier_class | binary | 0.1403109 | 5 | 0.001886026 | Preprocessor1_Model005 |
| 5 | 0.01000000 | roc_auc | binary | 0.9287602 | 5 | 0.005023629 | Preprocessor1_Model005 |
| 6 | 0.01000000 | accuracy | binary | 0.8503704 | 5 | 0.007300149 | Preprocessor1_Model006 |
| 6 | 0.01000000 | brier_class | binary | 0.1433502 | 5 | 0.002394401 | Preprocessor1_Model006 |
| 6 | 0.01000000 | roc_auc | binary | 0.9222099 | 5 | 0.006045245 | Preprocessor1_Model006 |
| 7 | 0.01000000 | accuracy | binary | 0.8525926 | 5 | 0.009633190 | Preprocessor1_Model007 |
| 7 | 0.01000000 | brier_class | binary | 0.1420321 | 5 | 0.003782513 | Preprocessor1_Model007 |
| 7 | 0.01000000 | roc_auc | binary | 0.9206950 | 5 | 0.007970750 | Preprocessor1_Model007 |
| 1 | 0.01623777 | accuracy | binary | 0.8188889 | 5 | 0.005629142 | Preprocessor1_Model008 |
| 1 | 0.01623777 | brier_class | binary | 0.1616034 | 5 | 0.002086724 | Preprocessor1_Model008 |
| 1 | 0.01623777 | roc_auc | binary | 0.8959839 | 5 | 0.006575368 | Preprocessor1_Model008 |
| 2 | 0.01623777 | accuracy | binary | 0.8362963 | 5 | 0.007444352 | Preprocessor1_Model009 |
| 2 | 0.01623777 | brier_class | binary | 0.1538296 | 5 | 0.001384560 | Preprocessor1_Model009 |
| 2 | 0.01623777 | roc_auc | binary | 0.9052567 | 5 | 0.002742090 | Preprocessor1_Model009 |
| 3 | 0.01623777 | accuracy | binary | 0.8362963 | 5 | 0.006186405 | Preprocessor1_Model010 |
| 3 | 0.01623777 | brier_class | binary | 0.1496267 | 5 | 0.002705010 | Preprocessor1_Model010 |
| 3 | 0.01623777 | roc_auc | binary | 0.9153666 | 5 | 0.007604319 | Preprocessor1_Model010 |
| ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ | ⋮ |
| 5 | 61.58482 | accuracy | binary | 0.7577778 | 5 | 0.0115410640 | Preprocessor1_Model131 |
| 5 | 61.58482 | brier_class | binary | 0.2138267 | 5 | 0.0011151672 | Preprocessor1_Model131 |
| 5 | 61.58482 | roc_auc | binary | 0.8449092 | 5 | 0.0061148461 | Preprocessor1_Model131 |
| 6 | 61.58482 | accuracy | binary | 0.7674074 | 5 | 0.0098444695 | Preprocessor1_Model132 |
| 6 | 61.58482 | brier_class | binary | 0.2127550 | 5 | 0.0010660400 | Preprocessor1_Model132 |
| 6 | 61.58482 | roc_auc | binary | 0.8453169 | 5 | 0.0060603434 | Preprocessor1_Model132 |
| 7 | 61.58482 | accuracy | binary | 0.7718519 | 5 | 0.0086741025 | Preprocessor1_Model133 |
| 7 | 61.58482 | brier_class | binary | 0.2120647 | 5 | 0.0010521195 | Preprocessor1_Model133 |
| 7 | 61.58482 | roc_auc | binary | 0.8455773 | 5 | 0.0060577024 | Preprocessor1_Model133 |
| 1 | 100.00000 | accuracy | binary | 0.6074074 | 5 | 0.0124363951 | Preprocessor1_Model134 |
| 1 | 100.00000 | brier_class | binary | 0.2368593 | 5 | 0.0009043861 | Preprocessor1_Model134 |
| 1 | 100.00000 | roc_auc | binary | 0.8310725 | 5 | 0.0056761244 | Preprocessor1_Model134 |
| 2 | 100.00000 | accuracy | binary | 0.6074074 | 5 | 0.0124363951 | Preprocessor1_Model135 |
| 2 | 100.00000 | brier_class | binary | 0.2357033 | 5 | 0.0009584880 | Preprocessor1_Model135 |
| 2 | 100.00000 | roc_auc | binary | 0.8297361 | 5 | 0.0056658875 | Preprocessor1_Model135 |
| 3 | 100.00000 | accuracy | binary | 0.6074074 | 5 | 0.0124363951 | Preprocessor1_Model136 |
| 3 | 100.00000 | brier_class | binary | 0.2357781 | 5 | 0.0009893254 | Preprocessor1_Model136 |
| 3 | 100.00000 | roc_auc | binary | 0.8286669 | 5 | 0.0056627580 | Preprocessor1_Model136 |
| 4 | 100.00000 | accuracy | binary | 0.6074074 | 5 | 0.0124363951 | Preprocessor1_Model137 |
| 4 | 100.00000 | brier_class | binary | 0.2357890 | 5 | 0.0012697318 | Preprocessor1_Model137 |
| 4 | 100.00000 | roc_auc | binary | 0.8284353 | 5 | 0.0058590361 | Preprocessor1_Model137 |
| 5 | 100.00000 | accuracy | binary | 0.6074074 | 5 | 0.0124363951 | Preprocessor1_Model138 |
| 5 | 100.00000 | brier_class | binary | 0.2347382 | 5 | 0.0011317313 | Preprocessor1_Model138 |
| 5 | 100.00000 | roc_auc | binary | 0.8288569 | 5 | 0.0056654048 | Preprocessor1_Model138 |
| 6 | 100.00000 | accuracy | binary | 0.6074074 | 5 | 0.0124363951 | Preprocessor1_Model139 |
| 6 | 100.00000 | brier_class | binary | 0.2345973 | 5 | 0.0011312843 | Preprocessor1_Model139 |
| 6 | 100.00000 | roc_auc | binary | 0.8286722 | 5 | 0.0056750661 | Preprocessor1_Model139 |
| 7 | 100.00000 | accuracy | binary | 0.6074074 | 5 | 0.0124363951 | Preprocessor1_Model140 |
| 7 | 100.00000 | brier_class | binary | 0.2343394 | 5 | 0.0011876321 | Preprocessor1_Model140 |
| 7 | 100.00000 | roc_auc | binary | 0.8287848 | 5 | 0.0056720982 | Preprocessor1_Model140 |
for(metric in unique(METRICS$.metric)){
metrics <- METRICS %>% filter(.metric==metric)
plot <- ggplot(metrics, aes(x=penalty, y=mean, color=as.factor(hidden_units))) + geom_line() + scale_x_log10() + labs(title=metric)
print(plot)
}
BEST <- select_best(RES, metric = 'accuracy')
BEST
| hidden_units | penalty | .config |
|---|---|---|
| <int> | <dbl> | <chr> |
| 7 | 0.1128838 | Preprocessor1_Model042 |
MODEL <- WF %>%
finalize_workflow(BEST) %>%
fit(TRAIN)
MODEL
══ Workflow [trained] ══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: mlp()
── Preprocessor ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
5 Recipe Steps
• step_normalize()
• step_dummy()
• step_nzv()
• step_corr()
• step_lincomb()
── Model ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
a 11-7-1 network with 92 weights
inputs: fixed.acidity volatile.acidity citric.acid residual.sugar free.sulfur.dioxide total.sulfur.dioxide density pH sulphates alcohol chlorides_Lots
output(s): ..y
options were - entropy fitting decay=0.1128838
Example: neural networks (regression)#
data(EX9.BIRTHWEIGHT, package='regclass')
# This is not usually right if want to investigate testing performances.
TRAIN <- EX9.BIRTHWEIGHT
REC <- recipe(Birthweight ~ ., TRAIN) %>%
step_normalize(all_numeric_predictors()) %>%
step_dummy(all_nominal_predictors()) %>%
step_nzv(all_predictors()) %>%
step_corr(all_predictors()) %>%
step_lincomb(all_predictors())
WF <- workflow() %>%
add_recipe(REC) %>%
add_model(mlp( mode = 'regression', hidden_units = tune(), penalty = tune() ))
GRID <- expand.grid( hidden_units = c(2,3), penalty = 10^seq(-2,1,length=15) )
RES <- WF %>%
tune_grid(
resamples = vfold_cv(TRAIN, v = 5),
grid = GRID,
)
METRICS <- collect_metrics(RES)
METRICS
→ A | warning: A correlation computation is required, but `estimate` is constant and has 0 standard deviation, resulting in a divide by 0 error. `NA` will be returned.
There were issues with some computations A: x1
There were issues with some computations A: x1
| hidden_units | penalty | .metric | .estimator | mean | n | std_err | .config |
|---|---|---|---|---|---|---|---|
| <dbl> | <dbl> | <chr> | <chr> | <dbl> | <int> | <dbl> | <chr> |
| 2 | 0.01000000 | rmse | standard | 500.34200416 | 5 | 27.78098862 | Preprocessor1_Model01 |
| 2 | 0.01000000 | rsq | standard | 0.11557329 | 5 | 0.05032802 | Preprocessor1_Model01 |
| 3 | 0.01000000 | rmse | standard | 514.41492660 | 5 | 25.03946676 | Preprocessor1_Model02 |
| 3 | 0.01000000 | rsq | standard | 0.07509909 | 5 | 0.02697816 | Preprocessor1_Model02 |
| 2 | 0.01637894 | rmse | standard | 492.35315438 | 5 | 19.92295040 | Preprocessor1_Model03 |
| 2 | 0.01637894 | rsq | standard | 0.12062614 | 5 | 0.02622690 | Preprocessor1_Model03 |
| 3 | 0.01637894 | rmse | standard | 491.98653197 | 5 | 18.37399980 | Preprocessor1_Model04 |
| 3 | 0.01637894 | rsq | standard | 0.12164642 | 5 | 0.03845223 | Preprocessor1_Model04 |
| 2 | 0.02682696 | rmse | standard | 516.37838731 | 5 | 28.15773717 | Preprocessor1_Model05 |
| 2 | 0.02682696 | rsq | standard | 0.07208546 | 4 | 0.03804960 | Preprocessor1_Model05 |
| 3 | 0.02682696 | rmse | standard | 505.25170670 | 5 | 21.36287174 | Preprocessor1_Model06 |
| 3 | 0.02682696 | rsq | standard | 0.08323415 | 5 | 0.04277127 | Preprocessor1_Model06 |
| 2 | 0.04393971 | rmse | standard | 525.74500034 | 5 | 19.65821297 | Preprocessor1_Model07 |
| 2 | 0.04393971 | rsq | standard | 0.04341909 | 5 | 0.02249748 | Preprocessor1_Model07 |
| 3 | 0.04393971 | rmse | standard | 510.49917381 | 5 | 22.08933728 | Preprocessor1_Model08 |
| 3 | 0.04393971 | rsq | standard | 0.09457031 | 4 | 0.01637357 | Preprocessor1_Model08 |
| 2 | 0.07196857 | rmse | standard | 490.09232893 | 5 | 20.78144716 | Preprocessor1_Model09 |
| 2 | 0.07196857 | rsq | standard | 0.13148280 | 5 | 0.02021711 | Preprocessor1_Model09 |
| 3 | 0.07196857 | rmse | standard | 475.98731310 | 5 | 24.07979661 | Preprocessor1_Model10 |
| 3 | 0.07196857 | rsq | standard | 0.17049092 | 5 | 0.04620822 | Preprocessor1_Model10 |
| 2 | 0.11787686 | rmse | standard | 471.04992567 | 5 | 14.09626010 | Preprocessor1_Model11 |
| 2 | 0.11787686 | rsq | standard | 0.18146804 | 5 | 0.02421311 | Preprocessor1_Model11 |
| 3 | 0.11787686 | rmse | standard | 502.03561725 | 5 | 16.16797090 | Preprocessor1_Model12 |
| 3 | 0.11787686 | rsq | standard | 0.11848598 | 5 | 0.02220887 | Preprocessor1_Model12 |
| 2 | 0.19306977 | rmse | standard | 514.19557350 | 5 | 20.74797506 | Preprocessor1_Model13 |
| 2 | 0.19306977 | rsq | standard | 0.05839087 | 5 | 0.01773080 | Preprocessor1_Model13 |
| 3 | 0.19306977 | rmse | standard | 505.10256344 | 5 | 24.47833230 | Preprocessor1_Model14 |
| 3 | 0.19306977 | rsq | standard | 0.11085628 | 5 | 0.02902798 | Preprocessor1_Model14 |
| 2 | 0.31622777 | rmse | standard | 510.32814056 | 5 | 22.04282333 | Preprocessor1_Model15 |
| 2 | 0.31622777 | rsq | standard | 0.08208713 | 5 | 0.03050509 | Preprocessor1_Model15 |
| 3 | 0.31622777 | rmse | standard | 497.31421593 | 5 | 14.03847181 | Preprocessor1_Model16 |
| 3 | 0.31622777 | rsq | standard | 0.09585007 | 5 | 0.03419205 | Preprocessor1_Model16 |
| 2 | 0.51794747 | rmse | standard | 501.06108491 | 5 | 15.59500375 | Preprocessor1_Model17 |
| 2 | 0.51794747 | rsq | standard | 0.10274716 | 5 | 0.02478217 | Preprocessor1_Model17 |
| 3 | 0.51794747 | rmse | standard | 509.04153544 | 5 | 22.07454587 | Preprocessor1_Model18 |
| 3 | 0.51794747 | rsq | standard | 0.09985624 | 5 | 0.03019326 | Preprocessor1_Model18 |
| 2 | 0.84834290 | rmse | standard | 493.68760683 | 5 | 17.52614032 | Preprocessor1_Model19 |
| 2 | 0.84834290 | rsq | standard | 0.10083736 | 5 | 0.02973864 | Preprocessor1_Model19 |
| 3 | 0.84834290 | rmse | standard | 498.82005941 | 5 | 8.34972820 | Preprocessor1_Model20 |
| 3 | 0.84834290 | rsq | standard | 0.14346320 | 5 | 0.02431361 | Preprocessor1_Model20 |
| 2 | 1.38949549 | rmse | standard | 504.56243534 | 5 | 14.14084938 | Preprocessor1_Model21 |
| 2 | 1.38949549 | rsq | standard | 0.09870968 | 5 | 0.02500284 | Preprocessor1_Model21 |
| 3 | 1.38949549 | rmse | standard | 489.76809898 | 5 | 14.97913350 | Preprocessor1_Model22 |
| 3 | 1.38949549 | rsq | standard | 0.11992775 | 5 | 0.02481531 | Preprocessor1_Model22 |
| 2 | 2.27584593 | rmse | standard | 491.91889212 | 5 | 20.71737596 | Preprocessor1_Model23 |
| 2 | 2.27584593 | rsq | standard | 0.11130928 | 5 | 0.04203917 | Preprocessor1_Model23 |
| 3 | 2.27584593 | rmse | standard | 486.16095784 | 5 | 22.57745993 | Preprocessor1_Model24 |
| 3 | 2.27584593 | rsq | standard | 0.15155921 | 5 | 0.03405783 | Preprocessor1_Model24 |
| 2 | 3.72759372 | rmse | standard | 496.04322669 | 5 | 18.10167754 | Preprocessor1_Model25 |
| 2 | 3.72759372 | rsq | standard | 0.11230776 | 5 | 0.01724078 | Preprocessor1_Model25 |
| 3 | 3.72759372 | rmse | standard | 513.14136223 | 5 | 20.32292991 | Preprocessor1_Model26 |
| 3 | 3.72759372 | rsq | standard | 0.10368869 | 5 | 0.02293471 | Preprocessor1_Model26 |
| 2 | 6.10540230 | rmse | standard | 534.69164622 | 5 | 10.94583540 | Preprocessor1_Model27 |
| 2 | 6.10540230 | rsq | standard | 0.07818785 | 5 | 0.01840885 | Preprocessor1_Model27 |
| 3 | 6.10540230 | rmse | standard | 515.85816570 | 5 | 15.29563700 | Preprocessor1_Model28 |
| 3 | 6.10540230 | rsq | standard | 0.06405564 | 5 | 0.02175738 | Preprocessor1_Model28 |
| 2 | 10.00000000 | rmse | standard | 521.08322173 | 5 | 24.08594502 | Preprocessor1_Model29 |
| 2 | 10.00000000 | rsq | standard | 0.08096775 | 5 | 0.01654549 | Preprocessor1_Model29 |
| 3 | 10.00000000 | rmse | standard | 521.26876409 | 5 | 22.82716363 | Preprocessor1_Model30 |
| 3 | 10.00000000 | rsq | standard | 0.09045355 | 5 | 0.02116048 | Preprocessor1_Model30 |
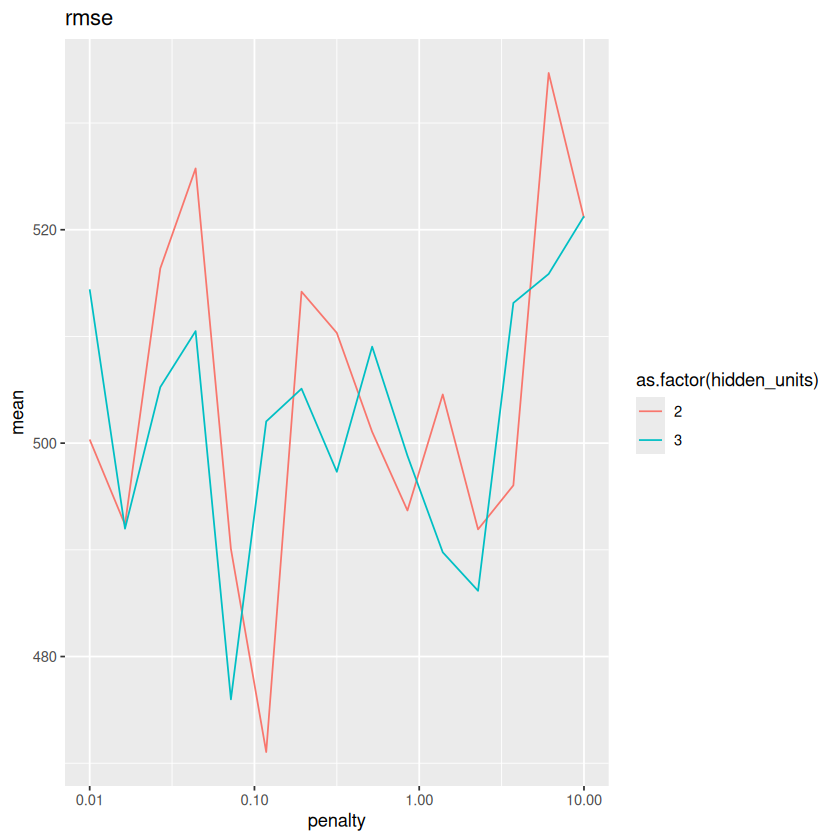
for(metric in unique(METRICS$.metric)){
metrics <- METRICS %>% filter(.metric==metric)
plot <- ggplot(metrics, aes(x=penalty, y=mean, color=as.factor(hidden_units))) + geom_line() + scale_x_log10() + labs(title=metric)
print(plot)
}
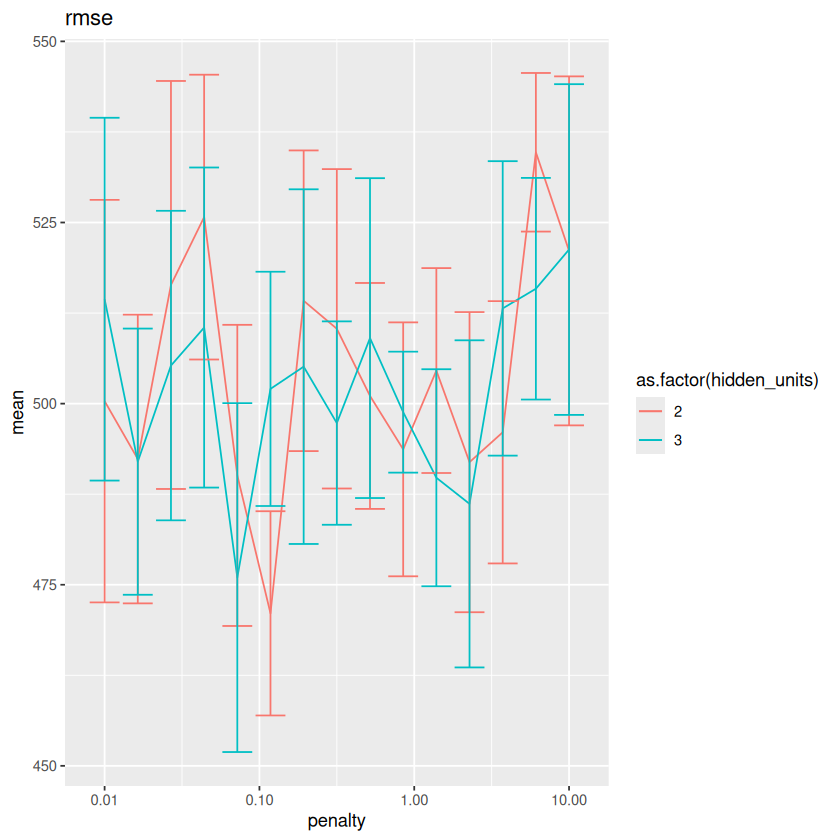
The results are very close to each other. What if we add \(std\_err\) as error bars?
for(metric in unique(METRICS$.metric)){
metrics <- METRICS %>% filter(.metric==metric)
plot <- ggplot(metrics, aes(x=penalty, y=mean, color=as.factor(hidden_units))) + geom_line() + scale_x_log10() + labs(title=metric)
plot <- plot + geom_errorbar(aes(ymin=mean-std_err, ymax=mean+std_err))
print(plot)
}
The error bars show that indeed the different models have very similar performances on this particular data. In the following, we still select the best model, but technically others are expected to work as well as the selected one.
BEST <- select_best(RES, metric = 'rmse')
BEST
| hidden_units | penalty | .config |
|---|---|---|
| <dbl> | <dbl> | <chr> |
| 2 | 0.1178769 | Preprocessor1_Model11 |
MODEL <- WF %>%
finalize_workflow(BEST) %>%
fit(TRAIN)
MODEL
══ Workflow [trained] ══════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════════
Preprocessor: Recipe
Model: mlp()
── Preprocessor ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
5 Recipe Steps
• step_normalize()
• step_dummy()
• step_nzv()
• step_corr()
• step_lincomb()
── Model ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
a 14-2-1 network with 33 weights
inputs: Gestation MotherAge MotherHeight MotherWeight FatherAge FatherHeight FatherWeight MotherEducation_College MotherEducation_HS FatherRace_Black FatherRace_White Father_Education_College Father_Education_HS Smoking_now
output(s): ..y
options were - linear output units decay=0.1178769
Pros and Cons#
Pros:
Can learn nonlinear relationships and create relevant predictors automatically through the use of hidden layers.
Does well when distributions resemble a Normal (symmetric, bell-shaped) curve (many image, text, or speech based problems that humans do) as long as there is a lot of data.
Once trained, predictions are very fast.
Deep learning, which are massive neural networks, actually are amazingly effective at what they are tuned to do.
Cons:
Overhyped early in its history and now. Neural networks will not solve every problem ever created.
Often doesn’t work the best for business problems where distributions do not resemble a Normal curve.
Hard to interpret (like most other models).
Computationally intensive to train and tune (like most other good models).
Well-known for neural network researchers: A neural network is the second best way to solve any problem. The best way is to actually understand the problem.